Blue Native PAGE (BN-PAGE) behind the scenes
Summary
TLDRThis video script delves into the intricate process of Blue Native PAGE, a technique used to separate protein complexes without denaturing them, preserving their natural structures and interactions. The narrator shares their hands-on experience, from preparing the gel to running the electrophoresis, highlighting the unique aspects of Blue Native PAGE, such as using Coomassie Blue dye to impart a negative charge on proteins for migration. Despite the non-informative results, the process's aesthetic appeal and the learning experience are emphasized, showcasing the experimental journey's value beyond mere outcomes. The script also touches on subsequent steps like staining and potential analyses, offering insights into the broader context of protein study techniques.
Takeaways
- 🔬 Blue Native PAGE (Polyacrylamide Gel Electrophoresis) is a technique used to separate protein complexes without denaturing them, allowing for the observation of multimer formations.
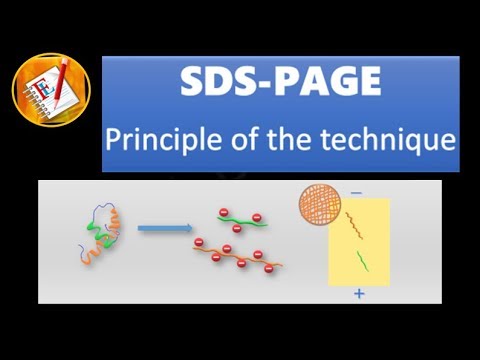
- 🧬 Unlike SDS PAGE, Blue Native PAGE does not unfold proteins but instead uses Coomassie Blue dye to impart a negative charge, enabling their movement through the gel based on size.
- 🔵 The Coomassie Blue dye gently binds to proteins and complexes without breaking them up, facilitating their migration through the gel.
- 🎨 The process results in visually striking blue gels, adding an aesthetic aspect to the scientific experiment.
- ⚡ The technique involves using electricity to move proteins through the gel, similar to SDS PAGE, but maintains protein complexes intact.
- 📊 Pre-cast gels with varying concentrations (e.g., 4-20%) and specialized ladders are utilized to accommodate the unique properties of proteins in Blue Native PAGE.
- 💧 Additional steps include careful sample preparation with glycerol to prevent samples from floating out of wells and ensuring a tight seal in the gel apparatus to avoid leaks.
- 🕒 After initial running, the inner buffer containing the dye is replaced with a fresh, undyed buffer to prevent over-dyeing and potential issues.
- 🔍 Post-run, proteins can be visualized directly if concentrated enough, or through staining techniques like Coomassie or silver staining for more sensitivity.
- 🧪 The experiment serves both an educational and exploratory purpose, demonstrating the application of Blue Native PAGE and providing hands-on experience with the technique.
Q & A
What is the primary purpose of using native PAGE over SDS PAGE?
-The primary purpose of using native PAGE over SDS PAGE is to separate protein complexes without unfolding the proteins, thereby preserving their native structure and allowing for the observation of multimers and protein complexes.
How does SDS PAGE differ from native PAGE in terms of protein treatment?
-SDS PAGE unfolds proteins and applies a uniform negative charge across them, facilitating their separation by size. Native PAGE, on the other hand, does not unfold proteins, allowing them to maintain their natural structures and complexes.
What is the role of the comassie blue dye in blue native PAGE?
-In blue native PAGE, comassie blue dye gently binds to proteins and their complexes without disrupting them, imparting a negative charge that enables their migration through the gel.
Why might a protein not migrate through a gel in SDS PAGE?
-A protein might not migrate through a gel in SDS PAGE if it does not have a negative charge under the conditions used, such as if the protein is inherently basic and positively charged, it won't be pulled through the gel without additional treatment.
What is the significance of replacing the dye with a fresh, undyed buffer during a blue native PAGE run?
-Replacing the dye with fresh, undyed buffer during a blue native PAGE run prevents over-dyeing and potential issues related to excessive staining, ensuring clearer results and better visualization of the proteins.
How does the presence of glycerol in the sample affect its migration in blue native PAGE?
-Glycerol is added to the samples to increase their density, ensuring they sink into the wells and do not float out, which is crucial for maintaining sample integrity during the electrophoresis process.
What is the purpose of using a special ladder in blue native PAGE?
-A special ladder is used in blue native PAGE because the normal ladders for SDS PAGE are denatured and would not behave correctly under native PAGE conditions. The special ladder is designed to migrate properly without denaturing.
What does the presenter mean by 'lazy staining' the gel, and why was it chosen?
-Lazy staining refers to a quick and simple staining method used by the presenter, chosen for convenience despite less informative results. It involves staining the gel directly, without prior destaining, to quickly visualize proteins.
What are the implications of spilling dye into the outer chamber of the gel apparatus?
-Spilling dye into the outer chamber can lead to unintended dye migration and contamination, affecting the clarity of the visualization and potentially the interpretation of the results, as it might obscure the visibility of the wells and migration lanes.
What are some post-run analyses that can be performed on proteins separated by blue native PAGE?
-Post-run analyses include staining the gel for visualization, conducting a second dimension PAGE for further separation, performing western blotting for specific protein identification, or analyzing the proteins via mass spectrometry for a comprehensive identification of the protein components.
Outlines

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowMindmap

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowKeywords

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowHighlights

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowTranscripts

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowBrowse More Related Video
5.0 / 5 (0 votes)