How to Calculate Adsorption Energy using Quantum ESPRESSO and DFT? [TUTORIAL]
Summary
TLDRIn this Phys Whiz tutorial, Manas Sharma demonstrates how to calculate the adsorption energy of a water molecule on the LiH (001) surface using Density Functional Theory (DFT) with Quantum ESPRESSO. The process involves three separate DFT calculations for the total system, the isolated molecule, and the surface. Sharma guides viewers through the use of BURAI, a GUI for Quantum ESPRESSO, to set up and run simulations, aiming to reproduce a study's adsorption energy value. The tutorial concludes with a comparison of the calculated energy to a reference value, highlighting the importance of pseudopotentials and energy cutoffs in achieving accurate results.
Takeaways
- 👨🏫 The tutorial is presented by Manas Sharma from Phys Whiz, focusing on calculating the adsorption energy of a molecule on a surface using density functional theory (DFT) and Quantum ESPRESSO.
- 💧 The specific molecule and surface discussed are H2O (water) and the LiH (001) surface, respectively.
- 📚 Adsorption energy is calculated using a formula that involves the total energy of the system and the energies of the isolated components.
- 📉 A negative adsorption energy indicates favorable adsorption, although some literature uses a reverse convention with a positive value indicating favorability.
- 📘 The tutorial references a 2017 paper and uses its supplementary material for the H2O-LiH system structure.
- 🧩 The structure provided includes various supercell sizes, but the tutorial uses the one with 16 Li and 16 H atoms, totaling 32 atoms in the surface.
- 📝 The structure is converted into a CIF file for use in Quantum ESPRESSO, as opposed to the original VASP format (POSCAR file).
- 🔬 The DFT calculation uses the PBE exchange-correlation functional aiming to reproduce the adsorption energy value reported in the paper.
- 🔄 Three separate calculations are needed: one for the total periodic system, one for the isolated H2O molecule, and one for the isolated LiH surface.
- 🛠️ BURAI, a GUI for Quantum ESPRESSO, is used to run the simulations and manage the project.
- 🔧 Pseudopotentials are carefully selected for each atom to ensure consistency and accuracy in the simulations.
- ⚙️ Energy cutoffs and charge cutoffs are set, with occupations fixed due to the likely semiconductor or insulator nature of the materials.
- 🔢 The tutorial demonstrates how to extract and calculate the final adsorption energy, comparing it to the value obtained in the referenced paper.
- 🔗 Links to the paper, simulation files, and a web app for unit conversion are provided in the description for further reference.
Q & A
What is the main topic of the tutorial presented by Manas Sharma in the Phys Whiz video?
-The main topic of the tutorial is calculating the adsorption energy of a molecule, specifically H2O on a LiH (001) surface, using density functional theory and Quantum ESPRESSO.
What is the general formula for calculating adsorption energy as described in the script?
-The general formula for calculating adsorption energy is the total DFT energy of the system minus the sum of the energies of the isolated molecule and the surface or slab.
What does a negative adsorption energy value indicate according to the convention used in the script?
-A negative adsorption energy value indicates that the adsorption is favorable, according to the convention used in the script.
What is the significance of the paper from 2017 mentioned in the script?
-The 2017 paper is significant because it provides the method and formula used for calculating the adsorption energy, and the script aims to reproduce the results from this paper using Quantum ESPRESSO.
How many atoms are in the surface structure used for the tutorial, and what is the total number of atoms including the H2O molecule?
-The surface structure used in the tutorial contains 32 atoms (16 Li and 16 H atoms), and including the H2O molecule, the total number of atoms is 35.
What is the purpose of using the same pseudopotentials for all atoms in the calculations?
-Using the same pseudopotentials for all atoms ensures consistency and accuracy in the calculations, preventing discrepancies that could arise from using different types of pseudopotentials.
Why is the energy cutoff for the wave function set to 50 Rydbergs and the charge cutoff to 500 Rydbergs in the calculations?
-The energy cutoff is set to 50 Rydbergs and the charge cutoff to 500 Rydbergs to ensure a balance between accuracy and computational speed, as a smaller cutoff speeds up simulations.
What is the role of BURAI in the calculations described in the script?
-BURAI is a GUI for Quantum ESPRESSO, and it is used to set up, run, and manage the SCF calculations for the total system, the isolated H2O molecule, and the isolated LiH surface.
What is the importance of ensuring that the water molecule does not interact with its neighboring periodic images in the isolated H2O calculation?
-Ensuring that the water molecule does not interact with its neighboring periodic images is important to accurately calculate the energy of an isolated molecule, avoiding any artificial interactions that could skew the results.
How does the script address the difference in conventions between a positive and a negative adsorption energy value?
-The script acknowledges that some papers use a reverse convention where a positive adsorption energy value indicates favorable adsorption. It clarifies that the results should be compared based on the magnitude, regardless of the sign.
What is the final calculated adsorption energy value in milli electron volts, and how does it compare to the value from the 2017 paper?
-The final calculated adsorption energy value is -223 milli electron volts, which is close to the value of 219 milli electron volts obtained in the 2017 paper, indicating good agreement despite slight differences.
What could be the reasons for the slight difference between the calculated adsorption energy and the value from the 2017 paper?
-The slight difference could be due to different choices of pseudopotentials or the energy cutoff not being fully converged, suggesting that a larger or smaller value might have been used in the paper.
Outlines

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowMindmap

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowKeywords

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowHighlights

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowTranscripts

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowBrowse More Related Video

Density Functional Theory (DFT)

Density Functional Theory, Part 2: Approximations

Measuring density of regular shaped, irregular shaped solid object, and liquid
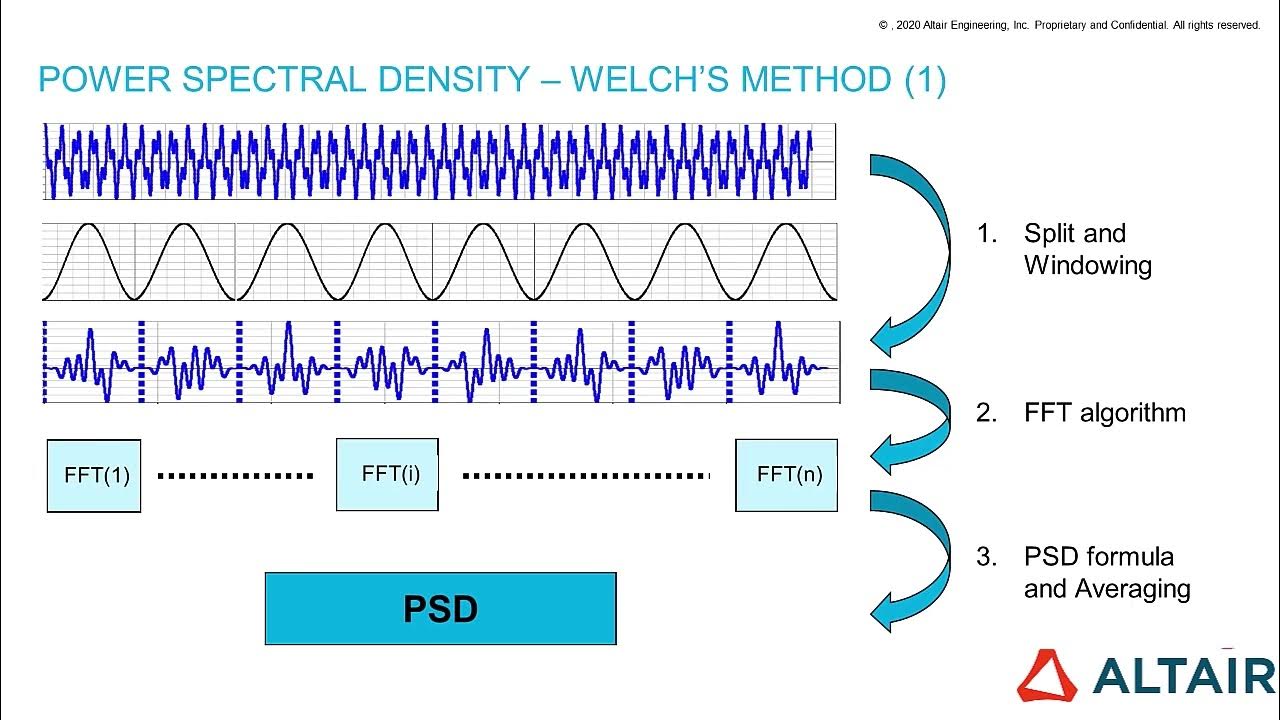
Altair Compose: Signal Processing - Power Spectral Density

MASSA JENIS (DENSITAS) - Menentukan Massa Jenis Suatu Zat | IPA Kelas 7

Why Quantum Algorithms? — Programming on Quantum Computers — Coding with Qiskit S2E1
5.0 / 5 (0 votes)