Gene Expression Analysis and DNA Microarray Assays
Summary
TLDRThis video script explores the fundamentals of gene expression analysis in biology, focusing on the genome's role in producing proteins. It delves into techniques like RT-PCR to understand gene expression in various contexts, including embryonic development and tissue-specific expression. The script highlights DNA microarray assays, a powerful tool for genome-wide expression studies, which can reveal gene alterations in diseases like cancer, offering insights for treatment development.
Takeaways
- 🧬 The key to understanding a living organism is to study the expression of its genome, which includes the identification of genes and the proteins they produce.
- 🔎 Gene expression analysis can reveal how different cell types within an organism perform and how gene expression might be altered in disease states like cancer.
- 📚 mRNA isolation is a fundamental step in analyzing gene expression, allowing for the identification of the proteins being produced by a cell.
- 🔬 Reverse transcriptase-polymerase chain reaction (RT-PCR) is a technique used to create a complementary DNA strand from an mRNA template, which is essential for further analysis.
- 🌟 The use of a poly-dT primer aids in initiating the reverse transcription process by binding to the poly-A tail of mRNAs.
- 🔄 After reverse transcription, DNA polymerase synthesizes the complementary DNA strand, resulting in double-stranded DNA known as cDNA.
- 👶 RT-PCR can be utilized to determine when a gene is being expressed during specific stages of embryonic development by comparing cDNA from different developmental stages.
- 📍 The technique can also identify which tissues within an organism are expressing a particular gene by extracting mRNA from various tissues and creating cDNA for analysis.
- 🌈 DNA microarray assays provide a comprehensive view of genome-wide expression by using a grid of DNA probes, each representing a specific gene, to which fluorescently labeled cDNA can hybridize.
- 🔍 By comparing the fluorescence patterns on a microarray, researchers can determine the expression levels of genes across different tissues or conditions, such as in cancer cells.
- 🛠 DNA microarray assays are a powerful tool in molecular biology, offering insights into gene expression alterations that can inform the understanding and treatment of diseases.
Q & A
What is the primary focus of studying living organisms in biology?
-The primary focus is to understand the expression of the organism's genome, which includes identifying the genes present and the proteins produced when these genes are expressed.
How does gene expression vary in different cell types within an organism?
-Gene expression varies depending on the cell type and its function within the organism. It can be studied by analyzing the mRNAs being made by a particular cell during transcription.
What is the purpose of isolating mRNAs from a cell?
-Isolating mRNAs allows researchers to perform techniques such as reverse transcriptase-polymerase chain reaction (RT-PCR) to understand gene expression patterns.
What is the role of reverse transcriptase in RT-PCR?
-Reverse transcriptase uses the mRNA as a template to generate a single-stranded DNA molecule that is complementary to the mRNA, effectively reversing the transcription process.
How does the poly-A tail of mRNAs assist in the RT-PCR process?
-The poly-A tail, a series of adenine RNA bases, allows the use of a poly-dT primer to initiate the reverse transcription process facilitated by reverse transcriptase.
What is the end product of the RT-PCR process?
-The end product is double-stranded DNA known as complementary DNA (cDNA), which is synthesized from the mRNA template.
How can RT-PCR be used to study gene expression during embryonic development?
-By isolating mRNAs from different stages of development and performing RT-PCR, researchers can identify cDNA fragments corresponding to mRNAs and determine when specific genes are being expressed.
What technique can be used to determine which tissues within an organism are expressing a particular gene?
-Extracting mRNA from different tissues and performing RT-PCR with amplification can reveal which tissue samples contain the mRNA associated with the gene of interest.
What is a DNA microarray assay and how does it work?
-A DNA microarray assay is a technique that uses a grid of spots, each containing many copies of a single-stranded DNA fragment (probe) representing a gene. cDNAs labeled with fluorescent tags are introduced to the array to hybridize with complementary DNA sequences, providing a visual representation of gene expression.
How can DNA microarray assays be used to understand genome-wide expression?
-By labeling cDNAs from different samples with different fluorescent colors and introducing them to the microarray, researchers can simultaneously obtain qualitative data on gene expression across various tissues or conditions.
What insights can DNA microarray assays provide into cancer cell gene expression?
-By comparing mRNAs from regular and cancer cells on the same microarray, researchers can identify genes that have been altered in cancer cells, which can lead to understanding the underlying mutations and suggesting new treatment approaches.
Outlines

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowMindmap

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowKeywords

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowHighlights

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowTranscripts

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowBrowse More Related Video

Prokaryotic gene structure

Gene Regulation and the Order of the Operon

TRYPTOPHAN OPERON- Easiest Explanation||Molecular Biology|TRP operon|Csirnet|gate|iitjam|msc|bsc|phd

Regulasi Ekspresi Gen #part1

Dogma sentral Biologi Molekuler dan Proses Ekspresi Genetik
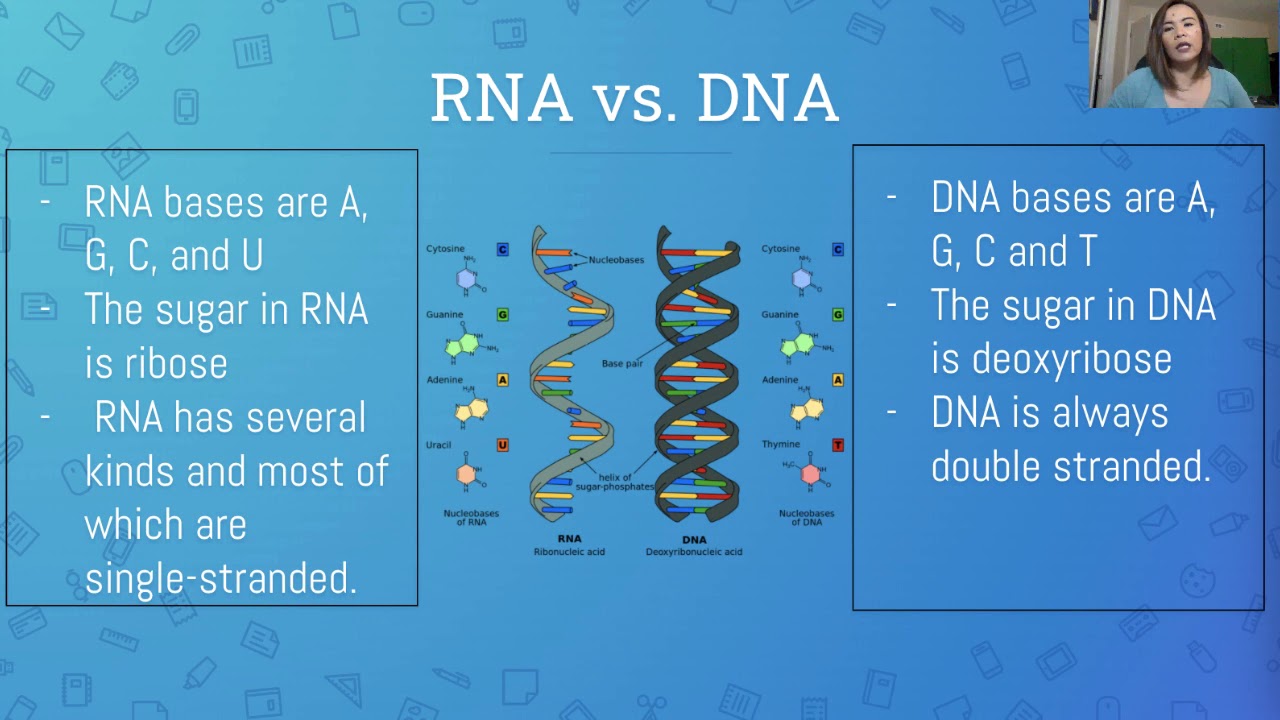
LESSON ON DNA, RNA and MUTATION | IN FILIPINO
5.0 / 5 (0 votes)