IB Biology A3.2 Classification & Cladistics
Summary
TLDRThis video script delves into the classification of living organisms, highlighting the shift from physical characteristics to DNA and genetic sequencing. It outlines the hierarchical system starting with domains to species, emphasizing the importance of common ancestry in grouping. The script also discusses the challenges of classification, such as the boundary paradox and the use of cladograms to represent evolutionary relationships. It concludes with the impact of DNA sequencing on reclassification, exemplified by the figwort family, and the addition of domains due to RNA sequencing.
Takeaways
- 🔬 Humans have a natural inclination to classify and label living organisms, which is crucial for understanding biodiversity.
- 🌐 The classification system in biology is hierarchical, starting broad with domains and becoming more specific through kingdoms, classes, orders, families, genera, and species.
- 🔬 Early classification was based on physical characteristics, but modern methods incorporate DNA and genetic sequencing to better understand evolutionary relationships.
- 🌿 There are three domains of life: Bacteria, Archaea, and Eukarya, which were established based on RNA sequencing.
- 📚 Taxonomy is the science of classifying organisms into groups, with domain being the broadest level of classification.
- 🧬 Genetic information, specifically DNA sequences, is considered the most reliable for classifying organisms and understanding their evolutionary origins.
- 🌱 Life is dynamic, with species changing and adapting to their environments, which can complicate classification efforts.
- 🌐 A cladogram is a diagram that represents the evolutionary relationships between different species based on shared characteristics.
- 🕰 The molecular clock is a concept that uses the rate of mutations to estimate the time since two species diverged from a common ancestor.
- 🔍 DNA sequencing has revolutionized the classification of species, leading to reclassifications as new evidence emerges.
- 🔄 The process of classification is not without challenges, as it involves subjective decisions and assumptions, such as the rate of mutations for the molecular clock.
Q & A
What is the primary reason humans classify and label living organisms?
-Humans classify and label living organisms to organize and manage information about them, enabling scientists and biologists to understand, record, and discuss species in a comprehensive and consistent manner.
What is the basis for the modern classification system in biology?
-The modern classification system in biology is based on a hierarchical structure that groups organisms according to their traits or evolutionary origins, with a shift towards using DNA and genetic sequencing.
What are the three domains that make up the largest groups of living organisms?
-The three domains that make up the largest groups of living organisms are Bacteria, Archaea, and Eukarya.
How does the hierarchical system of classification work?
-The hierarchical system of classification starts broad with the domain and becomes more specific through kingdom, class, order, family, genus, and species.
What is meant by the term 'taxonomic rank'?
-Taxonomic rank refers to the level of classification within the hierarchy, such as domain, kingdom, class, order, family, genus, and species.
What is the 'boundary paradox' in the context of biological classification?
-The 'boundary paradox' refers to the difficulty in determining when species have diverged enough to be considered separate species or different groups within the classification system.
Why is it important for classification to reflect evolutionary origins?
-Classification must reflect evolutionary origins because it allows for the sharing of traits between members of a group, enabling predictions to be made based on the classification, such as shared characteristics and adaptations.
What is a cladogram and how is it used in biological classification?
-A cladogram is a branching diagram that represents the evolutionary relationships among various groupings of organisms called clades, based on shared characteristics and common ancestry.
What is the molecular clock hypothesis and how is it used in classification?
-The molecular clock hypothesis is the idea that mutations occur at a consistent rate over time, allowing scientists to estimate the time since two species diverged from a common ancestor based on the differences in their DNA sequences.
How has DNA sequencing technology impacted the classification of species?
-DNA sequencing technology has greatly impacted the classification of species by providing more accurate information about evolutionary relationships, leading to reclassifications and a better understanding of species' ancestry.
What is the parsimony criterion in cladogram construction?
-The parsimony criterion is a method used in cladogram construction to find the simplest explanation for the evolutionary relationships among species, minimizing the number of evolutionary changes required.
Outlines

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowMindmap

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowKeywords

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowHighlights

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowTranscripts

This section is available to paid users only. Please upgrade to access this part.
Upgrade NowBrowse More Related Video

Manfaat dan Dasar Klasifikasi Makhluk Hidup

IB Biology A3.1 Diversity of Organisms

BIOLOGI SMA Kelas 12 - Materi Genetik | GIA Academy
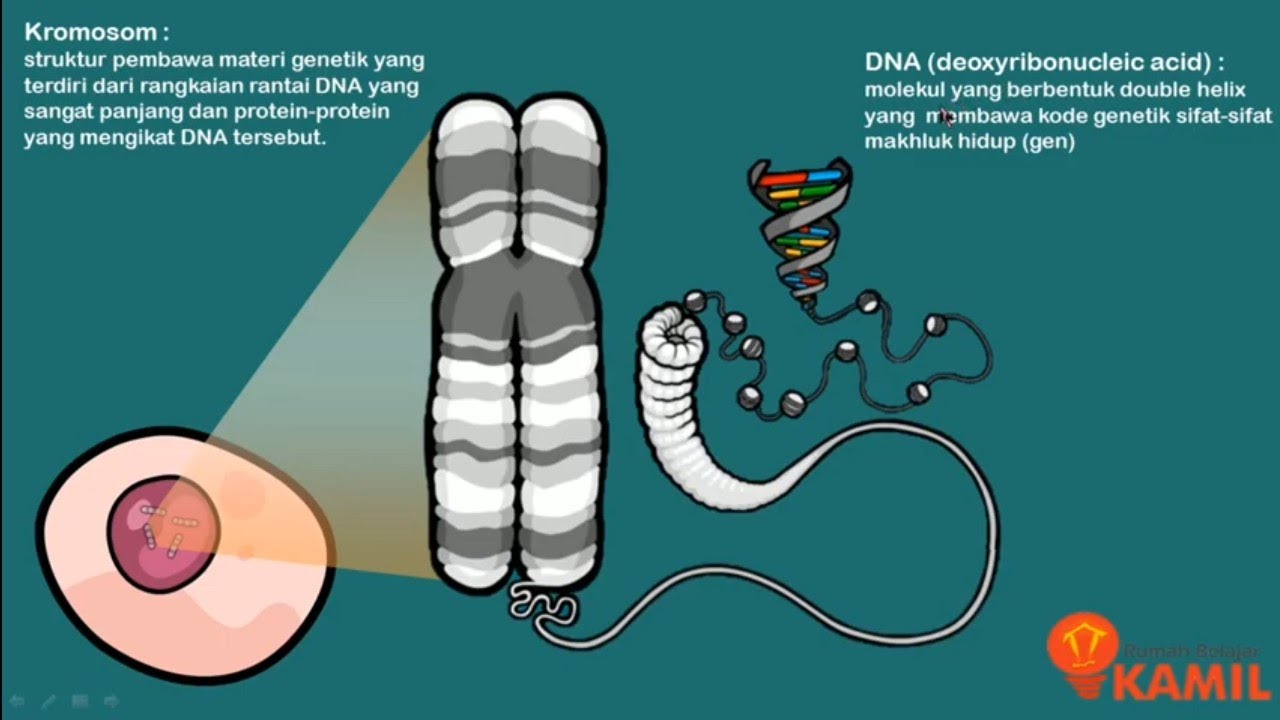
IPA Kelas 9 : Pewarisan Sifat I (Materi Genetik : Kromosom, DNA dan RNA)

How Are Organisms Classified? | Evolution | Biology | FuseSchool

The twisting tale of DNA - Judith Hauck
5.0 / 5 (0 votes)