t-SNE Simply Explained
Summary
TLDRThis video explores the T-Distributed Stochastic Neighbor Embedding (t-SNE), a powerful method in machine learning for reducing high-dimensional data to two or three dimensions for visualization. Unlike PCA, t-SNE focuses on preserving local structures by comparing data points pair-by-pair. The video delves into the algorithm's mechanics, including the use of a Gaussian distribution in high dimensions and a heavier-tailed distribution in lower dimensions to combat the curse of dimensionality. It concludes with a comparison to PCA, highlighting t-SNE's non-linearity and computational complexity.
Takeaways
- 📚 The video introduces a method called t-Distributed Stochastic Neighbor Embedding (t-SNE), used for reducing the dimensionality of high-dimensional datasets.
- 🔍 t-SNE is distinct from Principal Component Analysis (PCA) in that it operates on a pair-by-pair basis, focusing on preserving local relationships between data points.
- 📈 The method begins by defining the probability of one data point choosing another as its neighbor based on their distance in the high-dimensional space.
- 📉 t-SNE uses a function for calculating probabilities that resembles the division of two Gaussian probability density functions, aiming to maintain local structures in the lower-dimensional representation.
- 🌐 The algorithm dynamically selects a standard deviation (σ) for each data point to fit a certain number of neighboring data points within a specified number of standard deviations.
- 🔗 t-SNE creates a similarity score (p_ij) that represents the likelihood of data points i and j being grouped together, based on their closeness in the high-dimensional space.
- 📉 The goal of t-SNE is to find a lower-dimensional representation (y1, y2, ..., yn) that preserves the similarity (p_ij) between data points as computed in the high-dimensional space.
- 🔑 The challenge of dimensionality reduction is addressed by using a Student's t-distribution with a degree of freedom of one (a Cauchy distribution) in the lower-dimensional space, which has heavier tails than a Gaussian distribution.
- 📝 The algorithm employs the KL Divergence to measure the difference between the original high-dimensional similarities (p_ij) and the new low-dimensional similarities (q_ij), aiming to minimize this difference.
- 🤖 t-SNE uses gradient descent to adjust the positions of the lower-dimensional points (y_i, y_j) to minimize the KL Divergence, thus finding an optimal representation.
- 🚀 t-SNE is more computationally complex than PCA due to its focus on individual data point relationships and is particularly effective for datasets that are not linearly separable.
Q & A
What is the full name of the method discussed in the video?
-The full name of the method discussed in the video is T-Distributed Stochastic Neighbor Embedding, often abbreviated as t-SNE.
What is the primary goal of t-SNE?
-The primary goal of t-SNE is to reduce the dimensionality of a high-dimensional data set to a much lower-dimensional space, typically two or three dimensions, to enable visualization of the data.
How does t-SNE differ from Principal Component Analysis (PCA) in terms of its approach to dimensionality reduction?
-t-SNE operates on a pair-by-pair basis, focusing on preserving the local relationships between individual data points in the new lower-dimensional space, whereas PCA works on a global level, focusing on directions of maximum variance in the data set.
What is the significance of the probability P(J|I) in t-SNE?
-P(J|I) represents the probability that, given data point I, it would choose data point J as its neighbor in the original high-dimensional space. It is a key component in defining the local relationships between data points.
How does t-SNE address the 'curse of dimensionality' when mapping high-dimensional data to a lower-dimensional space?
-t-SNE addresses the curse of dimensionality by using a heavier-tailed distribution, specifically a Student's t-distribution with one degree of freedom (also known as a Cauchy distribution), to maintain higher probabilities for longer distances in the lower-dimensional space.
What role does the KL Divergence play in the t-SNE algorithm?
-The KL Divergence is used as a loss function in t-SNE to measure the difference between the original high-dimensional similarities (p_ij) and the new low-dimensional similarities (q_ij). The goal is to minimize the KL Divergence to ensure that the low-dimensional representation preserves the local structures of the high-dimensional data.
How does t-SNE handle the dynamic setting of the standard deviation σ_i for each data point?
-t-SNE dynamically sets the standard deviation σ_i for each data point based on the number of neighboring data points within a certain number of standard deviations of the Gaussian distribution. This allows t-SNE to adapt to different densities in the data, with lower σ_i for denser regions and higher σ_i for sparser regions.
What is the motivation behind using the Gaussian distribution to define probabilities in the high-dimensional space?
-The motivation behind using the Gaussian distribution is that it naturally incorporates the distance between data points into the probability calculation, with closer points having higher probabilities of being chosen as neighbors and more distant points having lower probabilities.
How does t-SNE ensure that the local structures between data points are preserved in the lower-dimensional space?
-t-SNE ensures the preservation of local structures by minimizing the KL Divergence between the original high-dimensional similarities and the new low-dimensional similarities, thus maintaining the relative closeness of data points that were close in the original space.
What computational method is used to find the low-dimensional representations (y_i) in t-SNE?
-Gradient descent is used to find the low-dimensional representations (y_i) in t-SNE. The algorithm iteratively adjusts the positions of the data points in the lower-dimensional space to minimize the KL Divergence between the original and new similarity measures.
Outlines

Этот раздел доступен только подписчикам платных тарифов. Пожалуйста, перейдите на платный тариф для доступа.
Перейти на платный тарифMindmap

Этот раздел доступен только подписчикам платных тарифов. Пожалуйста, перейдите на платный тариф для доступа.
Перейти на платный тарифKeywords

Этот раздел доступен только подписчикам платных тарифов. Пожалуйста, перейдите на платный тариф для доступа.
Перейти на платный тарифHighlights

Этот раздел доступен только подписчикам платных тарифов. Пожалуйста, перейдите на платный тариф для доступа.
Перейти на платный тарифTranscripts

Этот раздел доступен только подписчикам платных тарифов. Пожалуйста, перейдите на платный тариф для доступа.
Перейти на платный тарифПосмотреть больше похожих видео
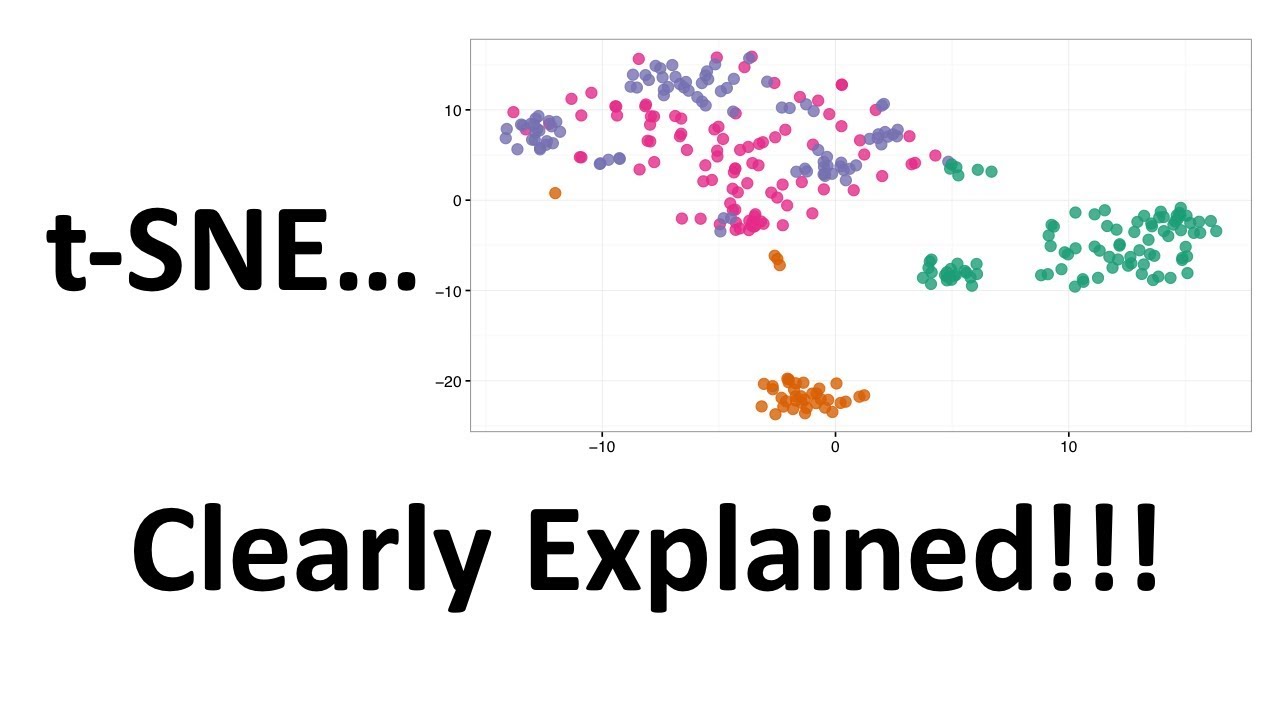
StatQuest: t-SNE, Clearly Explained

UMAP: Mathematical Details (clearly explained!!!)

Practical Intro to NLP 26: Theory - Data Visualization and Dimensionality Reduction

Bayesian Estimation in Machine Learning - Training and Uncertainties

Cara Uji Beda Dua Kelompok dengan Jamovi | Independent Sample t-test dan Paired Sample t-test
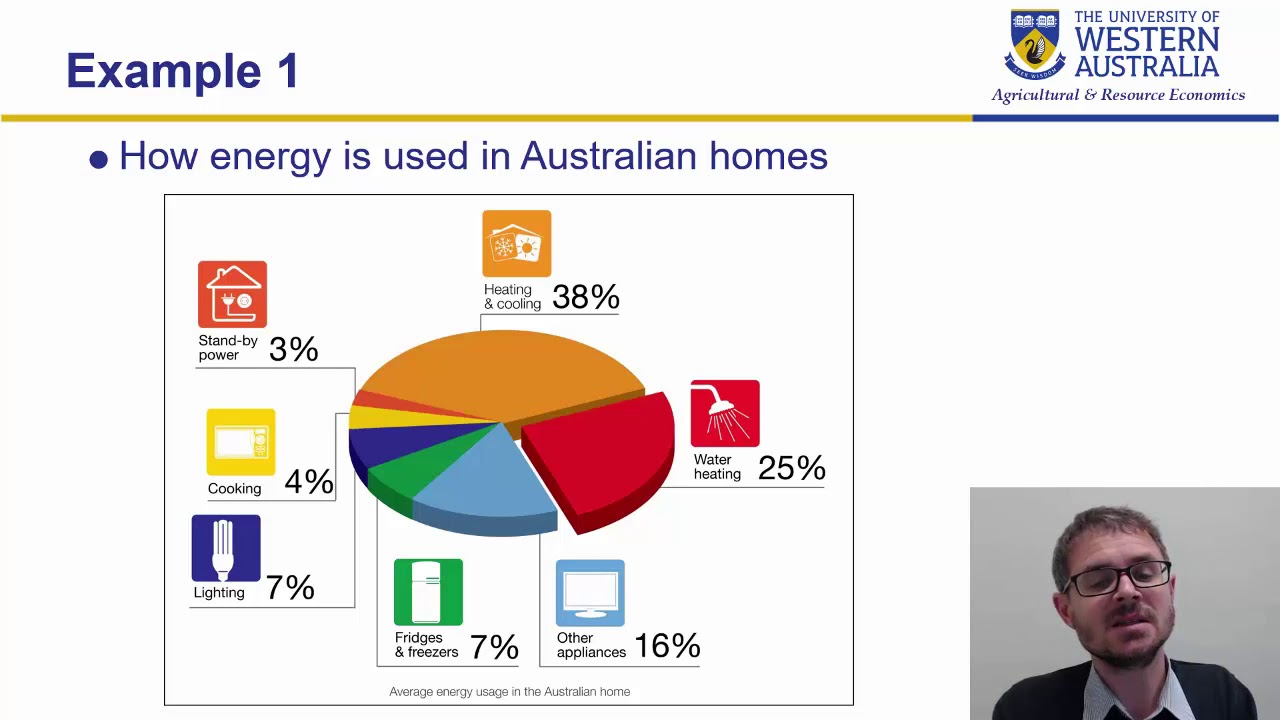
Wk 4 part5
5.0 / 5 (0 votes)
