Evolution 2023: Gene family expansions underlying a host-plant shift in Manduca sp. - Jay Goldberg
Summary
TLDRJay, a post-doctoral researcher, discusses his work on gene duplications in the Manduca genus, focusing on ecological co-evolution. He highlights the challenges of confirming co-evolution in plant-insect interactions and presents his findings on gene family changes in Manduca species, particularly involving glutathione transferases and odorant receptors. These genes may play roles in detoxification and host plant shifts. Jay aims to understand how natural selection shapes these interaction genes and their role in co-evolution, supported by extensive genome sequencing and analysis.
Takeaways
- 🔬 Jay's research focuses on gene duplications across the Manduka genus.
- 🌱 His broader research interest lies in co-evolution, which involves reciprocal selection during biological interactions.
- 🌍 Jay distinguishes between ecological co-evolution (species interactions in nature) and cellular/molecular co-evolution (molecular interactions within organisms).
- 🧬 Current research emphasizes ecological co-evolution, particularly in plant-insect interactions.
- 🦠 Microbial systems are often used to study co-evolution due to clear fitness effects and rapid generation times.
- 🐍 Jay mentions the debunking of the textbook example of garter snakes and toxic newts, highlighting the complexity of co-evolution.
- 🌼 His work involves studying the Manduka genus and their interactions with the Datura plant, noting the innate attraction of Manduka moths to Datura's floral volatiles.
- 📊 Jay uses whole genome sequencing and gene family analysis to study gene gain or loss events underlying host plant shifts.
- 🧪 Significant findings include the duplication of glutathione transferase genes in Manduka and extra copies of odorant receptor 67c in Manduka sexta.
- 🌐 The research aims to identify interaction genes mediating these co-evolutionary processes, contributing to the understanding of natural selection shaping these genes.
Q & A
What is the primary focus of Jay's post-doc research?
-Jay's post-doc research primarily focuses on gene duplications across the Manduca genus, with a broader emphasis on co-evolution, particularly ecological co-evolution.
What are the two types of co-evolution mentioned in the script?
-The two types of co-evolution mentioned are ecological co-evolution, which occurs when species interact in nature, and cellular or molecular co-evolution, which occurs when different molecules interact within an organism.
Why are microbial systems suitable for studying co-evolution?
-Microbial systems are suitable for studying co-evolution because they have clear fitness effects and rapid generation times, allowing for the observation of evolutionary changes in real time.
What challenge is associated with empirically assessing co-evolution in plant-insect interactions?
-Empirically assessing co-evolution in plant-insect interactions is challenging because there are many different interactions occurring, and any given trait of interest might be influenced by various environmental factors.
What recent findings have debunked a textbook example of co-evolution involving garter snakes and toxic newts?
-Recent findings have shown that the levels of toxins in newt populations are more closely related to their population structure than to the levels of resistance in their predators, indicating that selection is one-sided. The newts' toxins select for resistance in garter snakes, but the snakes' resistance does not select for higher toxin levels in the newts.
What is unique about the interaction between Datura plants and Manduca moths?
-The interaction between Datura plants and Manduca moths is unique because the floral volatiles of Datura plants are innately attractive to the moths, the larvae thrive on the toxic plants, and at least one species of Datura can be completely defoliated without any cost to seed production.
What method is Jay using to understand gene family changes in Manduca species?
-Jay is using whole genome sequencing and de novo assembly of Manduca species genomes, utilizing PacBio's HiFi reads for high accuracy and coverage.
What specific gene family did Jay's research find to have duplicated early in the evolution of the Manduca genus?
-Jay's research found that a glutathione transferase subfamily appears to have duplicated early in the evolution of the Manduca genus, with each species sampled having six to seven copies of this gene versus only one to two in the outgroups.
Why are the duplicated glutathione transferase genes interesting in the context of host plant colonization?
-The duplicated glutathione transferase genes are interesting because they are involved in detoxification of dietary compounds, which could have allowed Manduca species to colonize a diverse array of host plants by sub-functionalizing to detoxify different compounds.
How does Jay's research contribute to understanding co-evolution between Datura and Manduca?
-Jay's research aims to identify interaction genes mediating the interactions between Datura and Manduca, which will help understand how natural selection has shaped these genes and whether co-evolution has played a role in their adaptation.
Outlines

Cette section est réservée aux utilisateurs payants. Améliorez votre compte pour accéder à cette section.
Améliorer maintenantMindmap

Cette section est réservée aux utilisateurs payants. Améliorez votre compte pour accéder à cette section.
Améliorer maintenantKeywords

Cette section est réservée aux utilisateurs payants. Améliorez votre compte pour accéder à cette section.
Améliorer maintenantHighlights

Cette section est réservée aux utilisateurs payants. Améliorez votre compte pour accéder à cette section.
Améliorer maintenantTranscripts

Cette section est réservée aux utilisateurs payants. Améliorez votre compte pour accéder à cette section.
Améliorer maintenantVoir Plus de Vidéos Connexes

Polemik Baru Disertasi Bahlil, Jatam Ungkap Peran Peneliti UI

Bronfenbrenner's Ecological Systems: 5 Forces Impacting Our Lives

Restoration Blog: December 2021
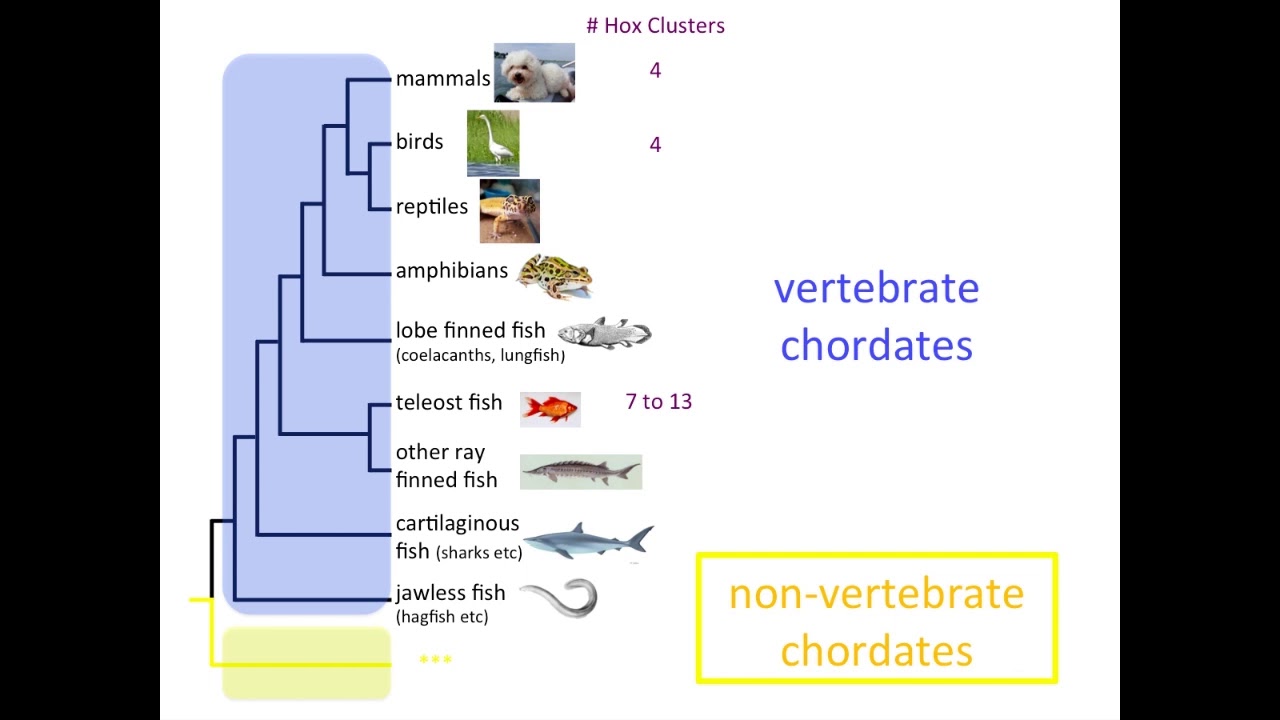
The Evolution of Hox Genes

L’evoluzione umana: gli inizi, 6 milioni di anni fa in Africa - Puntata 1

Billionaire Advice: The Benefits Of Operating Like An Entrepreneur, Even If You Aren't One | Forbes
5.0 / 5 (0 votes)
